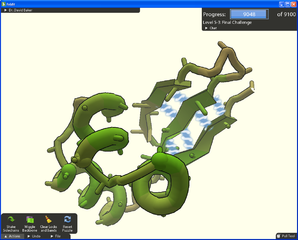
For 10 years, David Baker and his team of biochemists from the University of Washington had been trying to solve the crystal structure of a key protein in the development of HIV. The challenge was particularly difficult due to the extremely large number of possible outcomes.
They finally came up with the idea of stepping outside their usual research practice and creating an on-line game called Foldit. The concept? People spend hours playing computer games. Why not use that energy to solve real problems?
That was how 100,000 players started folding proteins. In September 2011, the 3D puzzle was solved. In just three weeks, the players had successfully produced an accurate model of the protein structure, providing new insights for the design of antiretroviral drugs. The project was a success not only due to the players’ energy but also to their different mindsets resulting in different approaches to finding solutions.
This is the first time that a game has been used to solve scientific problems: two major challenges have already been met using Foldit. What a wonderful achievement!
FIND OUT MORE:
- Video presentation of the game by the searchers and the players
- Publication dans Nature Structural and Molecular Biology
- Foldit, the game
FURTHER READING:
“Reinventing discovery, The New Era of Networked Science”, Michaël Nielsen, Princeton University Press
In our next rendez-vous, we’ll be taking you to Japan.